| |
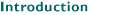

Brain
atlases are as essential for the research neuroscientist as are maps for
the geographer. Among the advantages of electronic atlases, compared with
the more conventional paper atlases, are that an electronic atlas can
contain images of all sections, allowing the brain to be "re-sectioned"
in any desired plane, can offer multiple levels of resolution and a range
of colors, can present structural features in 3D with variable transparency
of surface and internal components and free rotation so as to optimize
the user's view of structural relationships, can provide labels at the
whim of the user in more than one language, is readily edited and updated,
presents flexible indexing and ready access to other pertinent databases
and publications.
Why, then, are electronic
atlases not more readily available today? Part of the answer appears to
be that 1) until recently, working solutions had not been available with
desk top computers for image acquisition and display at histological levels
of resolution, 2) the drawing of structural boundaries by hand on a full
set of brain sections was too labor-intensive, and automated segmentation
was not yet feasible, given the relatively low-contrast inherent in stained
sections of brain tissue, 3) use of electronic atlases that display the
actual histological data sets was impractical because of limitations of
memory and band width, and 4) individual differences in brain form imposed
problems in defining a master ("canonical") brain for a given species.
A fuller presentation of these and related issues is available in Toga
AW, and Mazziotta JC. Brain Mapping: The Methods, San Diego: Academic
Press, 1996, 420 pages.
Sufficient computer power
is readily available now to resolve several of the problems summarized
above. Certain further problems touched on above still persist for specimens
as large and individually variable as the human brain, but have been solved
here in our project on the mouse brain. We have bypassed the issue of
individual variability by choosing a standard and widely used inbred strain
of mice named the C57BL/6 strain. Differences between strains can be turned
to advantage, and we predict that neuroscientists working with other inbred
strains will take advantage of differences they encounter between their
specimens and our "canonical" C57BL/6 mouse to explore the genetic basis
for individual differences in brain organization. The mouse is generally
recognized among biomedical scientists as a key mammal for research purposes,
and is clearly the mammal of choice for the increasingly dominant field
of genetic research. However, one may ask: is the mouse brain a good model
for the human brain? The mouse brain is obviously much smaller and simpler,
but it is of direct relevance with respect to human studies. The mouse
brain contains the same array of basic components laid out in a more linear
pattern, and serves very well as an introduction to mammalian brain architecture
in general.
|

