| |

The
general goals of the High Resolution Brain Atlas project are 1) to construct
atlases of the brain of the mouse, an animal that is becoming increasingly
important for neuroscience, particularly because of its position as the
mammal of choice in genetic research, and 2) to use the brain of this
small, relatively inexpensive and readily available animal as the test
case on which to work out issues of capture and integration of multidisciplinary
data, of 3D reconstruction from serial slices, and of data dissemination,
in the expectation that the lessons learned will become applicable directly
to the much larger normal and diseased human brain. The work accomplished
and the anticipated future directions are enumerated below.
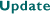

Creation of a 2D Digital
Atlas of the Mouse Brain
Our first goal, now essentially
achieved, has been to create a full 2D digital atlas of a two-month-old
female C57BL/6J mouse brain at 10 micron voxel resolution. The atlas is
based on a complete set of coronal sections stained alternately for cells
and for myelin, some of which were used in the paper atlas of Sidman,
Angevine and Taber Pierce (1971). The procedures for data capture, for
adjustment of staining intensity and hue among the sections, for alignment
of sections, and for nomenclature of gray and white matter structures
(annotations) are described here. For ease of calling up images over the
Internet at today's limited band width, the atlas has been divided arbitrarily
into eleven broad regions each of 1000 microns length-The 1st through
the olfactory bulb and the 11th through the caudal half of the cerebellum,
each region composed of 25 Nissl-stained sections paired with 25 myelin-stained
sections. A 12th region of 440 microns contains the caudalmost part of
the brain, the low medulla, and is composed of the final 11 Nissl-stained
sections paired with 11 myelin-stained sections. Also, the 2D atlas has
a 13th region containing as representative pair of Nissl- and myelin-stained
sections through each of the 33 spinal cord segments. For convenience
of access, a pair of sections at 200 micron intervals through the entire
brain is presented in the 2D atlas. In addition, composite images were
constructed by mapping a cell (Nissl-stained) image onto the adjacent
myelin image. An additional set of paired images is included with labels
and leader lines added ("annotated"), representing the coronal sections
at 200 micron intervals, plus additional sections that contain small structures
which lie entirely between the 200 micron increments. Thus, every named
structure can be visualized. The nomenclature matches that in the paper
atlas referred to above, with some additional or alternative names from
more recent publications. The user has the option of choosing English
or Latin nomenclature.
All histological processing
methods introduce distortions of brain tissue. For some purposes, such
as theoretical modelling, absolute measurement of cell and tissue dimensions,
strain comparisons and analysis of the actions of genes influencing strain
differences, and comparative neuroanatomy, it is important to warp the
histological images to some absolute "gold standard." The closest we come
today to a "gold standard" is a set of magnetic resonance images (MRI)
obtained over many hours at high magnetic strength, first on a fixed mouse
head and then on the same brain, carefully dissected from the skull. Absolute
dimensions are recorded, and all images are in alignment. Warping of our
histological images to an MRI series from brains of mice of the same strain,
age and sex are currently in progress. For other purposes, the unwarped
images may prove to be of greater value, particularly for the overlaying
of images obtained with other stains onto the "canonical" atlas's cell-
and myelin-stained images. Among the examples that come to mind are Golgi
and corresponding fluorescent images, immunohistochemical images, silver
degeneration and other axon-tracing images, in situ hybridizations
for gene expression, and pathological images. One could argue that a separate
atlas should be prepared by every common histological preparative method
(embedding in paraffin, glycol methacrylate, epoxy resin, or celloidin,
cryostat or vibratome sections, etc.), but as a practical compromise,
we propose to supplement the present 10 micron atlas of a celloidin-embedded
brain with 3 micron and 1 micron atlases based on differently processed
specimens, with and without image warping to the MRI standard (please
see below).
Mouse Brain 3D Atlas and
Volume Visualization
Another immediate goal is
to create a 3D voxel atlas of the C57BL/6J mouse brain from the present
2D section images. The critical factors are alignment of section images
and matching of stain intensities, because their precision controls the
efficiency and accuracy of 3D segmentation. The 10 micron images have
been aligned semiautomatically with newly written VoxelMath software,
followed by small further manual adjustments using pixel-based visual
landmarks. Full cell- and myelin-stained mouse brains have been constructed.
Further refinement of the alignment of serial images is anticipated, and
should precede final segmentation of gray and white matter structures.
Differences in staining intensity from section to section may be hardly
noticable when one studies sections in the standard manner, but the differences
become very important during 3D reconstruction. Precise matching of stain
intensities has proved to be unexpectedly difficult because, as we have
documented, the light, medium, and heavily stained tissue components do
not remap on the same linear scale. Not only do the positions of the stain
intensity histogram curves differ from section to section, but even their
shapes differ. Moderate success has been obtained by plotting the histogram
of voxel intensities in a relatively large 3D sample of brainstem, and
then mapping the entire series of section images to the shape and position
of that sample's histogram. We are working with texture mapping algorithms
to refine further the smoothing out of the staining differences, but even
now we are able to automate the 3D segmentation of selected gray and white
matter components, and expect to be adding the results to the atlas over
the coming months. The segmented volumes will then be connected via an
intuitively easy user interface to the informatics database of the High
Resolution Brain Atlas.
Data Dissemination
Our aim is that the images
be searchable through an informatics database by a query that addresses
either the images themselves or the names on a hierarchical structure
list. The database and the query range will be expanded gradually to link
other data from neuroanatomy, chemistry, pharmacology, physiology, psychology,
genetics and pathology. All images, including those in between the displayed
images at 200 micron intervals, will be distributed via this website on
demand by the user. We are setting up procedures and protocols to accomplish
the delivery, and ask any user with an immediate need for some subset
of the section images, or for atlas images resectioned electronically
at an orientation matching the user's own specimen(s), to send mail to
Richard
L. Sidman.


From research of the
past ~100 years, it is clear that maps are needed at different resolutions
to provide different kinds of information. Many aspects of structure delineation
in gray and white matter regions are best done at 10 micron voxel resolution,
whereas cell classification and texture-based segmentation appear to be
best accomplished at 3 microns, axon trajectories at 1 micron, and synaptic
organization in, say, retinal plexiform layers, glomeruli in cerebellum
and elsewhere, and in cortical sensory barrels and columns are resolvable
only by electron microscopy at < 0.1 micron. The map we propose to develop
at 1 micron resolution will meet the objective we and others have contemplated,
of plotting for the first time the detailed trajectories of white matter
tracts throughout the brainÑeven, for the larger caliber axons, down to
the level of individual myelinated fibers. Such a refined anatomy in the
mouse brain, especially cortico-cortical white matter anatomy, will likely
prove to be extrapolatable to other mammals, and should delineate the
fundamental white matter organization in even the human cerebrum. Such
a body of information is essential for interpreting human data obtained
by fMRI and related methods.
To plot such detailed anatomy
manually would be an immense undertaking. We anticipate that automated
and accurate segmentation will be feasible with thinner section images
captured at high resolution. It will become realistic to trace trajectories
of the larger myelinated axons for long distances through the brain, and
to store data on how constant are the relationships among axons over long
distances, especially in systems where we know from physiology that topographic
representation is preserved across synapses. Cell size, shape and packing
density, automatically recorded in the segmented 3D gray matter "clouds,"
should provide new useful and objective criteria for classification of
structural units in the brain in supplementation of input/output relationships
and chemical/genetic properties. The more complete the neuroanatomical
delineation of features in the mouse brain, the better will neuroscientists
be able to map new and old research data onto the atlas templates.
|

